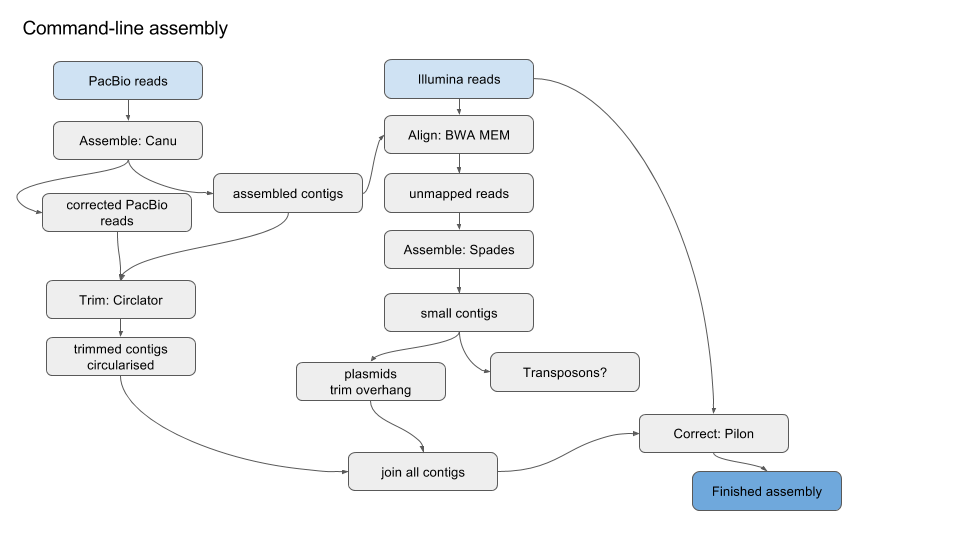
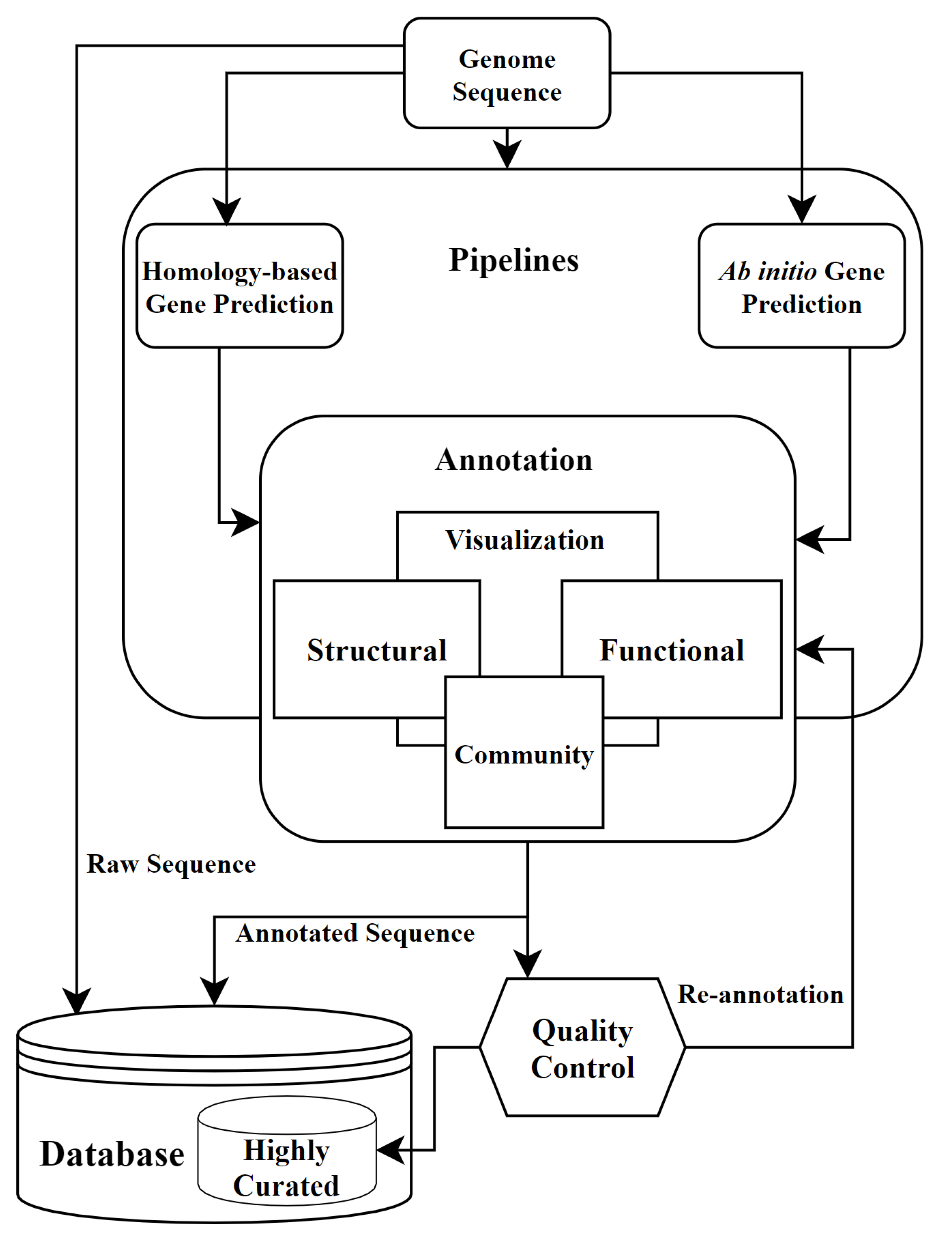
The bulk of the sequencing generated by the YCGA is whole exome sequencing, with whole genome sequencing as a small but growing portion. Whole-Exome and Whole-Genome Sequencing Analysis
#How to use novo screen annotation software
This page contains tips, advice and software tool descriptions for people who want to use the cluster for their analyses, as well as our current best practice procedures for WES and WGS sequencing (with RNA-seq, ChIP-seq and others coming soon). A description of the cluster and more information can be found on the Yale Center for Computing Resources pages. Yale has a number of high-performance compute clusters, and ruddle is the new cluster that will soon be available for the analysis of YCGA data.
